Impurity (Gini) measure implements binary decision trees and the three impurity measures or splitting criteria commonly used in binary decision trees are Gini impurity (IG), entropy (IH) and classification error (IE).

Contents
ToggleGini impurity
Used by the CART algorithm (tree classification and regression) for classification trees, the Gini impurity is a measure of how often a randomly chosen item in the set would be incorrectly labeled if it were randomly labeled based on the distribution of labels in the sub-set. together.
Mathematically, we can write the Gini impurity as follows:
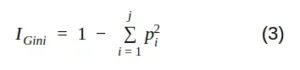
where j is the number of classes present in the node and p is the class distribution in the node.
Simple simulation with a heart disease dataset consisting of 303 rows and 13 attributes. The target includes 138 0 values and 165 1 values
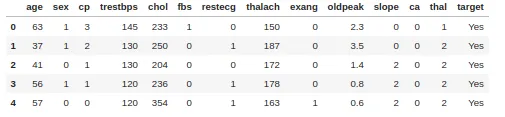
In order to create a decision tree from the dataset and determine which separation is best, we need a way to measure and compare the impurity in each attribute. The lowest impurity value in the first iteration will be the root node. we can write equation 3 in the form:
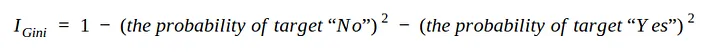
In this simulation, use only the sex, fbs (fasting blood glucose), exang (exercise-induced angina), and target attributes.
How to measure impurity in the Sex attribute:

- Left knot = 0.29
- Right knot = 0.49
Now that we have measured the impurity for both leaf nodes. We can calculate the total impurity with the weight average. The left node represented 138 patients while the right node represented 165 patients.
Total Gini impurity — Leaf node

We proceed in the same way with the other attributes:
- I_fbs_left = 0.268; right = 0.234; I_fbs = 0.249
- I_exang_left =0.596; right = 0.234; I_Exang = 0.399
Fbs (fasting blood sugar) has the lowest Gini impurity, so use it at the root node.
Gini impurity in quantitative data
Like weight which is one of the attributes to determine heart disease, for example we have the weight attribute:
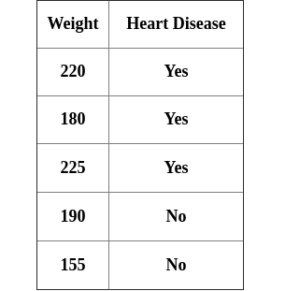

After ordering in ascending order, do the consecutive even-average and calculate the impurity for each value.
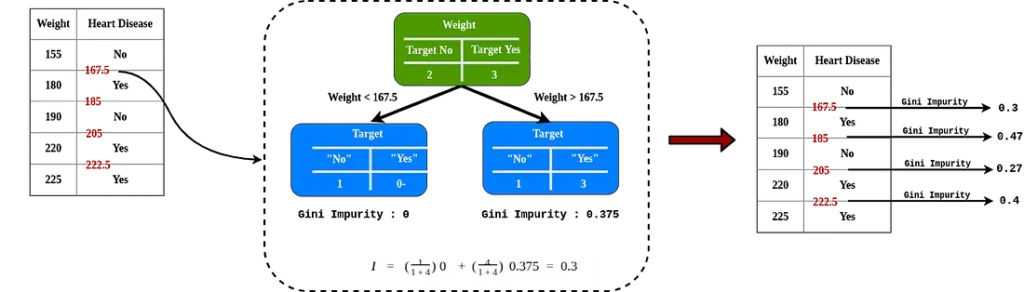
The lowest Gini impurity is weight < 205, this is the threshold value and the impurity value if used when we compare with another attribute.
For calculation reasons, it is also possible to do by quantile.
Gini impurity in qualitative data
We have a preferred color attribute for determining a person's gender:
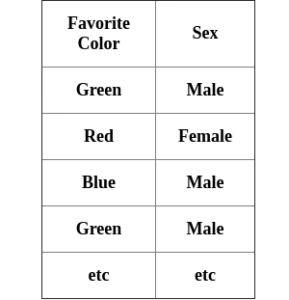

In order to know how impurity this attribute is, calculate an impurity score for each (as a Boolean value) as well as each possible combination.
Entropy
Used by the ID3, C4.5 and C5.0 tree generation algorithms. The information gain is based on the notion of entropy, the measure of entropy is defined as:
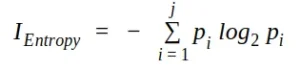
where j is the number of classes present in the node and p is the class distribution in the node.
Given the same case and data set, we need a way to measure and compare the entropy in each attribute. The highest entropy value in the first iteration will be the root node.
First we need to calculate the entropy in the Target attribute = 0.994.
We use the same split for sex:
- sex = 0 has for entropy 0.666
- sex = 1 has entropy 0.988
Now that we have measured the entropy for the two leaf nodes. We take the average of the weights to calculate the total entropy value.
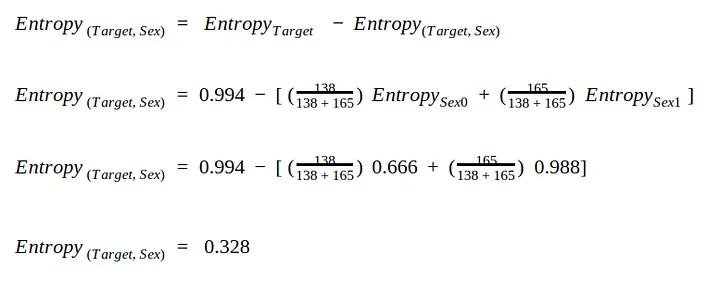

Entropy for column Fbs = 0.389
Entropy for column Exang = 0.224
Fbs (fasting blood sugar) has the highest entropy, so we will use it at the root node, exactly the same results we got with the Gini impurity.
Classification error / Misclassification impurity
Another measure of impurity is misclassification impurity or misclassification error. Mathematically, we can write the misclassification impurity as follows:
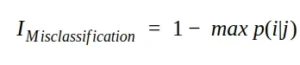
In terms of qualitative performance, this index is not the best choice because it is not particularly sensitive to different probability distributions (which can easily drive selection towards subdivision using Gini or entropy).
