Contents
ToggleAnswer Key Exercises on Exploratory Data Analysis
This page presents two corrected and detailed exercises with python code on exploratory data analysis.

Exercise 1
Consider a random sample of finishers from the New York City Marathon in 2002. This dataset is in the UsingR package. Load the library, then load the nym.2002 dataset.
library(dplyr)
data(name.2002, package="UsingR")Use boxplots and histograms to compare male and female finish times. Which of the summaries best describes the difference?
data(name.2002, package="UsingR")
male <- nym.2002 %>% filter(gender == 'Male')
female <- nym.2002 %>% filter(gender == 'Female')
mypar(1,2)
history(female$time, xlim= vs(100,600))
history(male$time, xlim= vs(100,600))
Both histograms have a similar distribution (skewed to the right). But the center of the histogram seems to be different. We can verify this by calculating the absolute difference between the mean and the median.
abs(mean(male$time) - mean(female$time))## [1] 23.11574abs(median(male$time) - median(female$time))## [1] 21.70833There is a difference of about 21-23 minutes between men and women. Males and females have similar right-skewed distributions, with the first 20 minutes shifted to the left.
Use dplyr to create two new data frames: male and female, with data for each gender. For men, what is the Pearson correlation between age and time to complete?
stud(male$age, male$time, hand = 'male')
horn(male$age, male$time)## [1] 0.2432273For women, what is Pearson's correlation between age and time to complete?
stud(female$age female$time, hand = 'female')
horn(female$age female$time)## [1] 0.2443156If we interpret these correlations without visualizing the data, we would conclude that the older we get, the slower we run marathons, regardless of gender. Examine the scatterplots and boxplots of hours stratified by age groups (20-24, 25-30, etc.). After looking at the data, what is the most reasonable conclusion?
groups_m <- split(male$time, floor(male$age/5)*5) # 10-14, 15-19, etc.
groups_f <- split(female$time, floor(female$age/5)*5) # 10-14, 15-19, etc.
mypar(1,2)
boxplot(groups_m)
boxplot(groups_f)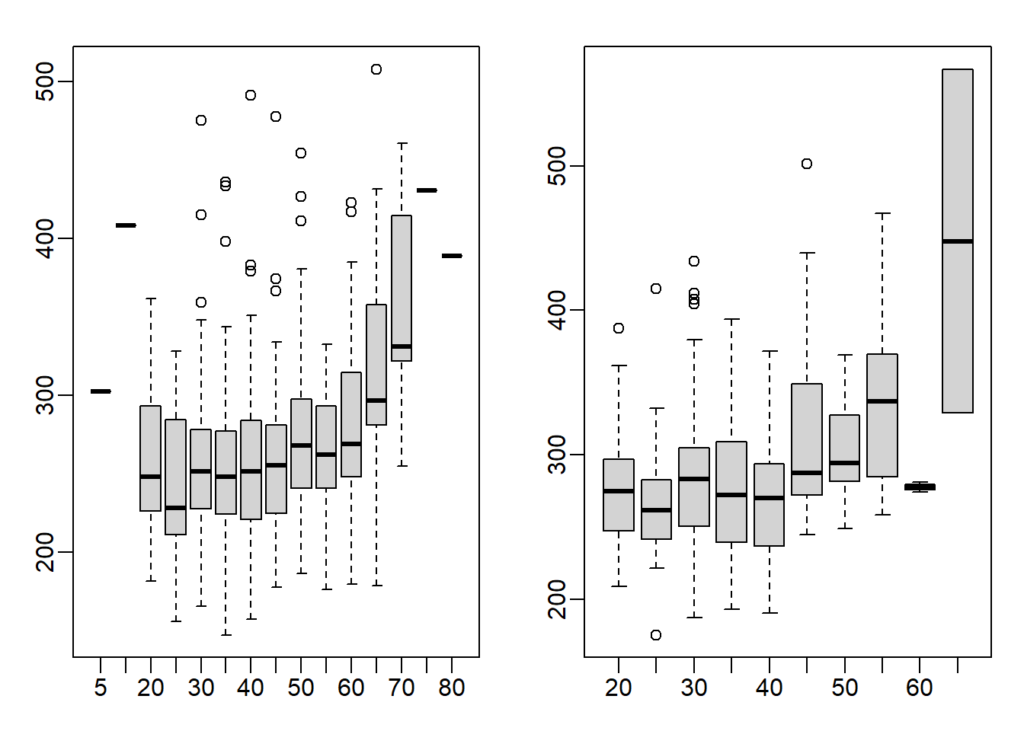
This is a tricky question because the question asks you to stratify the data into groups. Layering can be achieved via the split function. In order for each group to have a range of 5 (ex. 25-30), all age numbers will need to be rounded up or down so that the resulting numbers are divisible by 5. I rounded the numbers down down using the floor function.
Therefore, 40 represents the 40-44 age range. You can also use the ceiling function to stratify the data, which will then be rounded. So, 45 represents 41-45 age group. In the example below, the age of 42 is classified using the floor and ceiling functions.
floor(42/5)*5 ## [1] 40ceiling(42/5)*5## [1] 45Finish times are consistent until about our 40s, then we get slower.
Exercise 2
Let's load the data:
data(Chick Weight)
mypar()
stud(ChickWeight$Time, ChickWeight$weight, col=ChickWeight$Diet)
chick = reshape(ChickWeight, idvar=vs("Chick","Diet"), timevar="Time",
direction="wide")
chick = na.omit(chick)Focus on chick weights on day 4 (check chick column names and note numbers). How much does the average chick weight on day 4 increase if we add an outlier of 3000 grams? Specifically, what is the average day 4 chick weight, including the outlier chick, divided by the average day 4 chick weight without the outlier. Hint: use c to add a number to a vector.
chick_w4 <- chick[,'weight.4']
chick_w4_add <- append(chick_w4, 3000)
# or use function c
# chick_w4_add <- c(chick_w4, 3000)
chick_w4_add ## [1] 59 58 55 56 48 59 57 59 52 63 56 53 62
## [14] 61 55 54 62 64 61 58 62 57 58 58 59 59
## [27] 62 65 63 63 64 61 56 61 61 66 66 63 69
## [40] 61 62 66 62 64 67 3000mean(chick_w4_add) - mean(chick_w4) # Difference between with and without outlier## [1] 63.90966mean(chick_w4_add)/mean(chick_w4) # Ratio between with and without outlier## [1] 2.062407In the first part, we saw how sensitive the mean is to outliers. Now let's see what happens when we use the median instead of the mean. Calculate the same ratio, but now using the median instead of the mean. Specifically, what is the median day 4 chick weight, including the outlier chick, divided by the median day 4 chick weight without the outlier.
median(chick_w4_add) - median(chick_w4) # difference## [1] 0median(chick_w4_add)/median(chick_w4) # ratio## [1] 1Now try the same with the standard deviation example (the sd function in R). Add a chick weighing 3000 grams to the weights of day 4 chicks. How much does the standard deviation change? What is the standard deviation with the outlier chick divided by the standard deviation without the outlier chick?
sd(chick_w4_add) - sd(chick_w4) # difference## [1] 429.1973sd(chick_w4_add)/ sd(chick_w4) # ratio## [1] 101.2859Compare the result above to the median absolute deviation of R, which is calculated with the mad function. Note that the mad is not affected by adding a single outlier. The mad function in R includes the scaling factor 1.4826, so mad and sd are very similar for a sample from a normal distribution. What is the MAD with the outlier chick divided by the MAD without the outlier chick?
crazy(chick_w4_add) - crazy(chick_w4) # difference## [1] 0crazy(chick_w4_add)/ crazy(chick_w4) # ratio## [1] 1Our final question is about how the Pearson correlation is affected by an outlier from the Spearman correlation. The Pearson correlation between x and y is given in R by cor(x,y). The Spearman correlation is given by cor(x,y,method=”spearman”).
Plot the weights of day 4 and day 21 chicks. We can see that there is a general trend, with lower weight chicks on day 4 having low weight again on day 21, and the same for lower weight chicks raised.
Calculate the Pearson correlation of the chick weights from day 4 and day 21. Now calculate how much the Pearson correlation changes if we add a chick that weighs 3000 on day 4 and 3000 on day 21. Again divide the correlation of Pearson with the chick outlier on the calculated Pearson correlation without the outliers.
chick_w21 <- chick[, 'weight.21']
chick_w21## [1] 205 215 202 157 223 157 305 98 124 175 205 96 266 142 157 117
## [17] 331 167 175 74 265 251 192 233 309 150 256 305 147 341 373 220
## [33] 178 290 272 321 204 281 200 196 238 205 322 237 264stud(chick_w4, chick_w21)
horn(chick_w4,chick_w21) # correlation before## [1] 0.4159499chick_w21_add <- append(chick_w21, 3000)
horn(chick_w4_add, chick_w21_add) # correlation after outlier## [1] 0.9861002horn(chick_w4_add, chick_w21_add)/horn(chick_w4,chick_w21) # ratio between after and before## [1] 2.370719Record the chick weights on day 4 of diet 1 as the x vector. Record the chick weights on day 4 of diet 4 as a vector y. Perform a t-test comparing x and y (in R, the function t.test(x,y) will perform the test). Then perform a Wilcoxon test of x and y (in R the function wilcox.test(x,y) will perform the test). A warning will appear stating that an exact p-value cannot be calculated with links, so an approximation is used, which is fine for our purposes.
Perform a t-test of x and y, after adding a single 200 gram chick to x (the chicks in diet 1). What is the p-value of this test? The p-value of a test is available with the following code: t.test(x,y)$p.value
x <- chick %>% filter(Diet == 1)
x <- x[,'weight.4']
y <- chick %>% filter(Diet == 4)
y <- y[,'weight.4']
t.test(x,y)$p.value # t.test result with no outlier## [1] 7.320259e-06wilcox.test(x,y)$p.value # wilcox result with no outlier## Warning in wilcox.test.default(x, y): cannot compute exact p-value ## with ties## [1] 0.0002011939x_add <- vs(x,200) # outlier added
t.test(x_add,y)$p.value # t-test after outlier## [1] 0.9380347Do the same for the Wilcoxon test. The Wilcoxon test is robust to outliers. Also, it has fewer assumptions than the t-test about the distribution of the underlying data.
wilcox.test(x_add,y)$p.value # even with outlier, p-value is not perturbed## Warning in wilcox.test.default(x_add, y): cannot compute exact p- ## value with ties## [1] 0.0009840921We will now study a possible drawback of the Wilcoxon-Mann-Whitney test statistic. Using the following code to create three boxplots, showing the true diet weights 1 versus 4, then two modified versions: one with an additional 10 gram difference and one with an additional 100 gram difference. Use the x's and y's as defined above, NOT the ones with the outlier added.
library(rafalib)
mypar(1,3)
boxplot(x,y)
boxplot(x,y+10)
boxplot(x,y+100)
What is the difference in the t-test statistic (obtained by t.test(x,y)$statistic) between adding 10 and adding 100 to all values in group y? Take the t-test statistic with x and y + 10 and subtract the t-test statistic with x and y +100. The value must be positive.
t.test(x,y+10)$statistics - t.test(x,y+100)$statistics ## t ## 67.75097Examine the Wilcoxon test statistic for x and y+10 and for x and y+100. Since the Wilcoxon works on ranks, once the two groups show complete separation, i.e. all points in group y are greater than all points in group x, the statistic will not change , regardless of the magnitude of the difference. Similarly, the p value has a minimum value, regardless of the distance between the groups.
This means that the Wilcoxon test can be considered less powerful than the t-test in some contexts. In fact, for small samples, the value of p cannot be very small, even when the difference is very large. What is the value of p if we compare c(1,2,3) to c(4,5,6) using a Wilcoxon test?
wilcox.test(x,y+10)$p.value## Warning in wilcox.test.default(x, y + 10): cannot compute exact p- ## value with ties## [1] 5.032073e-05wilcox.test(x,y+100)$p.value## Warning in wilcox.test.default(x, y + 100): cannot compute exact p- ## value with ties## [1] 5.032073e-05wilcox.test(vs(1,2,3),vs(4,5,6))$p.value # Answer## [1] 0.1What is the value of p if we compare c(1,2,3) to c(400 500 600) using a Wilcoxon test?
wilcox.test(vs(1,2,3),vs(400,500,600))$p.value## [1] 0.1