Contents
ToggleCART regression and classification: use of the decision tree
Here is a tutorial for implementing CART Regression and CART Classification.
As has been explained, decision trees are the non-parametric supervised learning approach. In addition to classification with continuous target data, we also often find cases with discrete target data called regression. In regression, the simplest way may be to use linear regression to solve this case. This time the way to solve the regression case will use a tree decision.

CART and Least Squares for Regression
For regression trees, two common impurity measures are:
Least squares. This method is similar to minimization least squares in a linear model. The divisions are chosen to minimize the residual sum of squares between the observation and the mean in each node.
The smallest absolute deviations. This method minimizes the average absolute deviation from the median within a node. The advantage of this method over least squares is that it is not as sensitive to outliers and provides a more robust model. The disadvantage is insensitivity when dealing with data sets containing a large proportion of zeros.
CART in classification cases uses Gini Impurity in the process of splitting the dataset into a decision tree. On the other hand, CART in regression cases uses least squares, splits are chosen intuitively to minimize the residual sum of squares between the observation and the mean in each node. In order to know the “best” distribution, we must minimize the RSS:

Slicing and optimization
This simulation uses a “dummy” dataset as follows

As mentioned before, in order to know the “best” distribution, we need to minimize the RSS. First, we calculate RSS by dividing it into two regions, starting with index 0
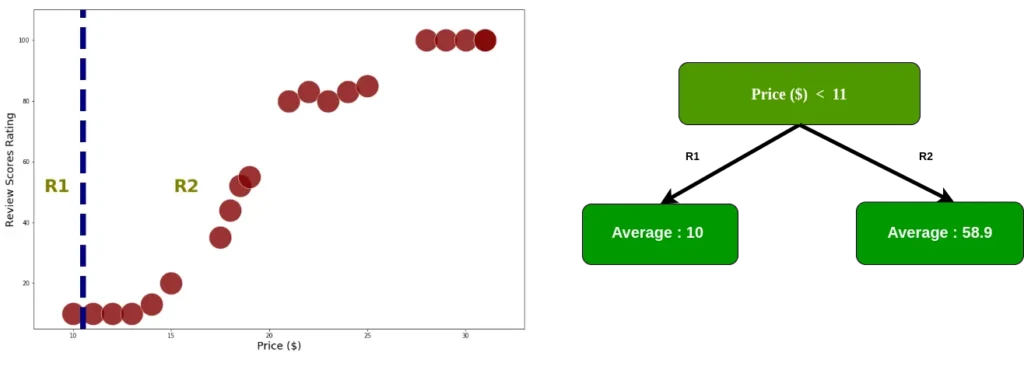
Since the data is already divided into two regions, we add the residual square for each index data. Moreover, we calculate RSS each node using equation 2.0
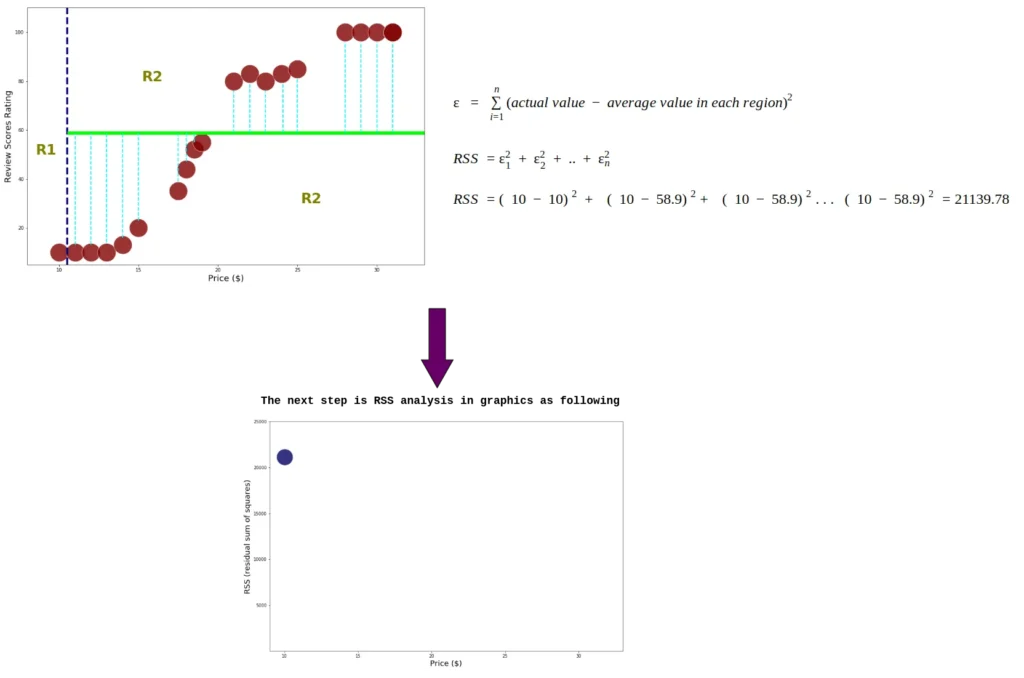
This process continues until the RSS is calculated in the last index:

The price with threshold 19 has smallest RSS, in R1 there are 10 data in price < 19, so we will split the data in R1. In order to avoid overfitting, we set the minimum data for each region >= 6. If the region contains less than 6 data, the division process in that region stops.
Split data with threshold 19

calculate RSS in R1, the process in this section is the same as the previous process, performed only for R1. It is possible to make as many branches as you wish, be careful to keep a consistent mass of points!

Which gives the following regression tree after 2 branches:

With multiple columns
Simply calculate the RSS for each predicator and root/branch the one with the smallest RSS (be careful to use preprocessing and an adequate metric in case of unbalanced data!)
CART for classification
For classification we use theimpurity by Gini. As an example, we take a heart disease dataset with 303 rows and 13 attributes. The target includes 138 0 values and 165 1 values
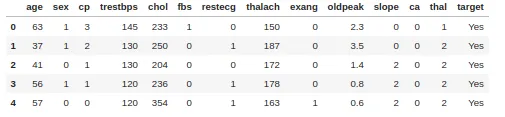
Let's calculate the Gini impurity for the Sex column


For Fbs we have 0.360. For Exang 0.381. Fbs has the smallest Gini impurity, so this will be our root.
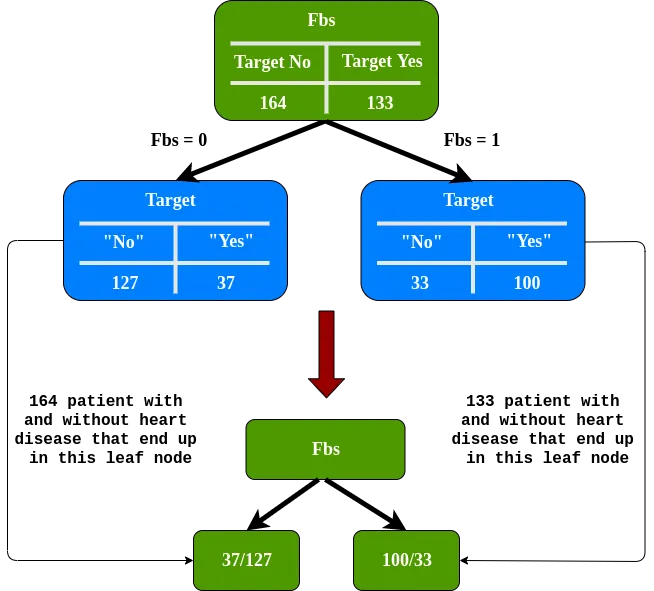
we need to determine to what extent Sex and Exang separate these patients in the left node of Fbs
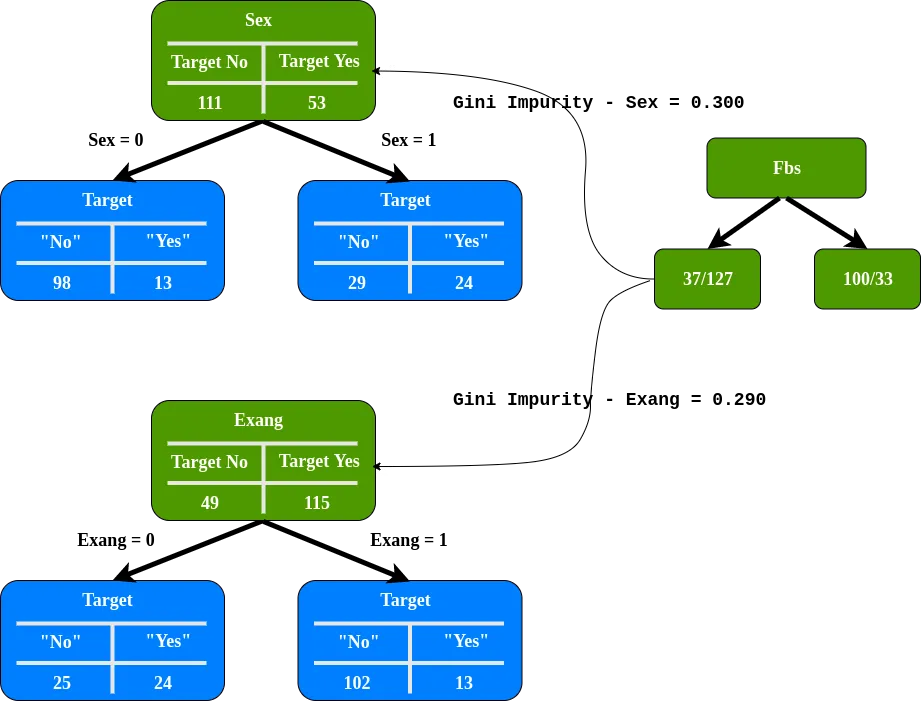
Exang (exercise-induced angina) has the lowest Gini impurity, we will use it at this node to separate patients.
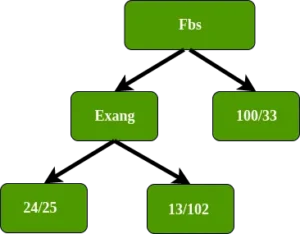
In the left node of Exang, how far does this separate these 49 patients (24 with heart disease and 25 without heart disease. Since only the sex attribute remains, we put the sex attribute in the left node of Exang Exang.
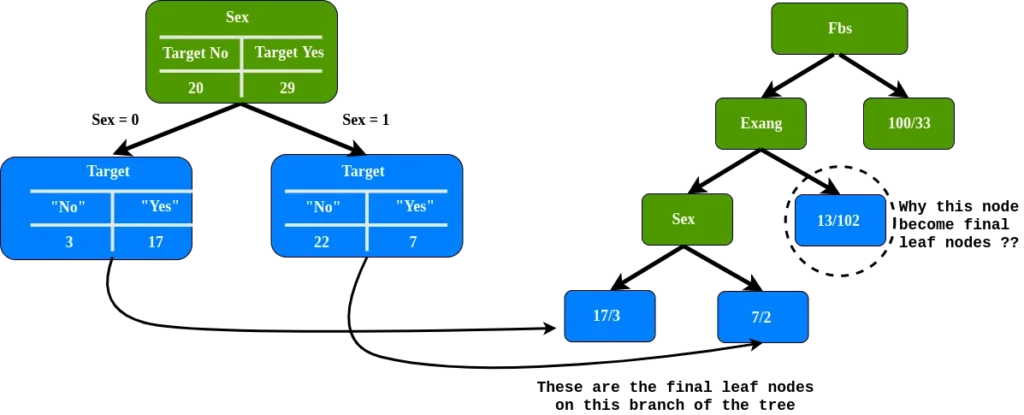
As we can see, we have final leaf nodes on this branch, but why is the leaf node circled, including the final node?
Note: the circled leaf node, 89% do not have heart disease
Do these new sheets separate patients better than before?
In order to answer these questions, we need to compare the Gini impurity using the sex attribute and the Gini impurity before using the sex attribute to separate patients.

The Gini impurity before using gender to separate patients is the lowest, so we do not separate this node using gender. The final leaf node on this branch of the tree. We do the same for the right branch of the root, which gives the following decision tree:
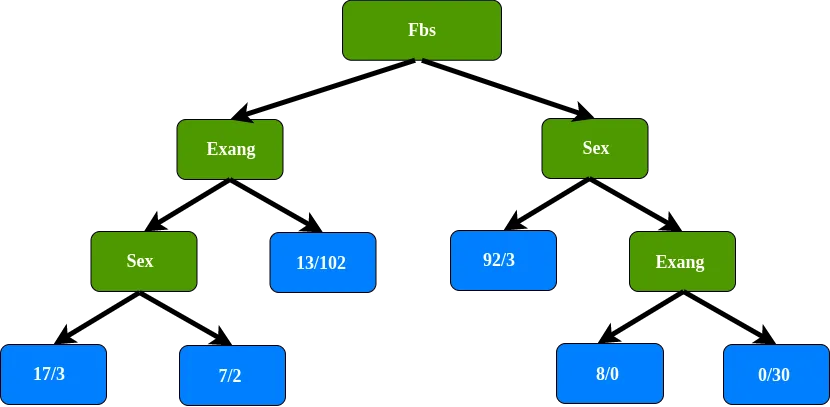
Classification recap
Main point during the data set splitting process
1. calculate all Gini impurity score
2. compare the Gini impurity score, after n before using a new attribute to separate the data. If the node itself has the lowest score, there is no point in separating the data
3. If data separation results in improvement, choose the separation with the lowest impurity score.
